Project under PMRF:
Structure-based thermodynamics of CRISPR/Cas9 genome editing: Insight from computer simulations
|
CRISPR/Cas9 system is a popular genome-editing tool that can transform any genome by removing, adding, or altering sections of the DNA sequence at a specific location. CRISPR/Cas9 is an RNA-dependent endonuclease, where a guide RNA (sgRNA) recognizes the DNA sequence based on RNA:DNA complementarity, and the Cas9 protein catalyzes the double-stranded break in the DNA at both target strand (tDNA containing sequence complementary to sgRNA) and the non-target strand (ntDNA) of the gene of interest.
Recently near-atomic resolution structures at various stages (apo-Cas9, Cas9:sgRNA, Cas9:sgRNA:tDNA, Cas9:sgRNA:tDNA: incomplete ntDNA; Cas9:sgRNA:dsDNA pre-catalytic, Cas9:sgRNA:dsDNA post-catalytic) of the CRISPR/Cas9 DNA editing pathway have been reported. Despite the advancement of structural and biochemical studies related to CRISPR/Cas9, we are far away from understanding the underlying structure-based free energy landscape associated with the editing pathway. The reported structures provide an excellent template for computational analysis. Using the experimentally determined structures as a template, I want to estimate the relative binding affinity (ΔΔG) of Cas9 in complex with the RNA:DNA hybrid upon DNA or RNA mutations by employing classical Molecular Dynamics (MD) simulation in an explicit water box to the template structures. Estimated energetics (ΔΔG) will then be linked to 3D structures. This methodology would be used to study the effect of Cas9 mutation on nucleotide binding with Cas9. The effect of Cas9 mutations in the energetics of RNA:DNA binding would be explored to understand which mutations favor or disfavor the off-target effect. Despite Cas9's massive potential in genome editing, off-target DNA cleavage by the CRISPR/Cas9 is another important concern that certainly limits its safety in therapeutic use. This necessitates understanding the detailed molecular mechanism of off-target effect and hypothesising novel Cas9 mutants with reduced off-target effects. Understanding the thermodynamics (ΔΔG) of the off-target effects in terms of atomic structures will provide a rationale for Cas9 mutant with reduced off-target effects. Additionally, Cas9’s constraints to cleave only in the presence of a specific PAM sequence limit its applicability over a wide range of genes. This necessitates a deeper understanding of the detailed molecular mechanism of how Cas9 recognizes specific nucleotides. Several key Cas9 residues are crucial in recognizing specific nucleotide residues and its mutations alter the nucleotide readability of the Cas9 protein. However, the molecular mechanism and associated thermodynamics behind how these mutations alter nucleotide selectivity are still unknown. MD simulation is an excellent complement to the experiments and helps establish a direct link between thermodynamics, kinetics, and molecular structures. |
|
Key Objectives: 1. Structure-based MD simulation of Cas9 at various stages of genome editing. 2. Effect of Cas9 mutation on the nucleotide binding affinity. 3. Energetics of PAM selectivity by Cas9. 4. Establish a bridge between the estimated energetics and molecular structures. |
Some critical regions of interest in Cas9 endonuclease
|
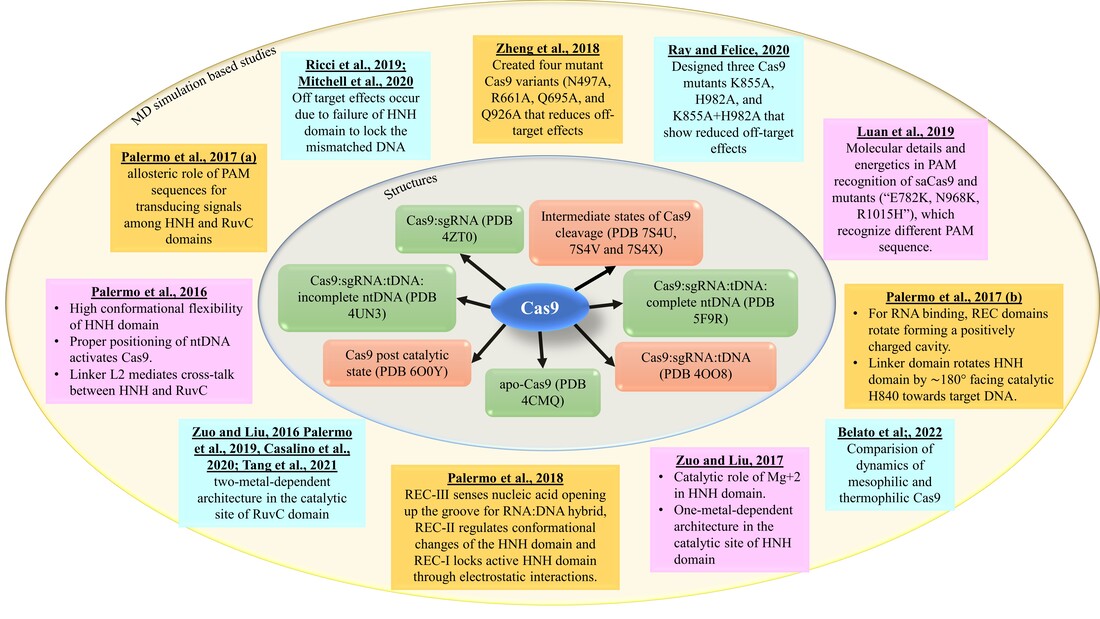
Literature survey:
The progress of structural and MD simulation-based studies on CRISPR/Cas9 system